Once again it is time for a BIG DATA download, which is useful when you need dozens or hundreds of scenes. People have come up with some Python (e.g., sentinelsat or Sentinel_download) and R libraries (e.g., sentinel2) in recent years to automatically copy Sentinel data from the ESA servers. All of these libraries are based on two Application Program Interfaces (APIs) provided by the ESA Data Hub used for browsing and downloading the data stored in the rolling archives.
We will take a closer look at ESA’s underlying, official script and we will use it to automatically download ALL records of any of your search queries. The script is named dhusget.sh and already deposited in our VM (or available for download here). You need to have a Linux Environment with wget installed (such as our VM) in order to use the script, which is a Unix shell script (.sh).
First of all, let us have a look at a blueprint of the command, we will adapt to your needs in the following:
/home/student/Documents/dhusget.sh -d https://scihub.copernicus.eu/dhus -l 100 -P 1 -C /media/sf_exchange/product_list.csv -u user123 -p password123 -o product -O /media/sf_exchange/sentineldata -R /media/sf_exchange/sentinel_data/failed.txt -F 'query'
Explanation: That is a lot of options! Each option is separated from the next by a blank. Anyway, you do not need to modify most of those options:
- /home/student/Documents/dhusget.sh: location of shell script in the VM
- -d https://scihub.copernicus.eu/dhus: the URL of the Data Hub Service
- -l 100: number of results per page (100 max.)
- -P 1: page number (each page contains a maximum of 100 results)
- -C /media/sf_exchange/product_list.csv: write all products found to a list and save it as a CSV text file
- -u user123: insert your ESA SciHUB username here
- -p password123: insert your ESA SciHUB password here
- -o product: activate download for the Product ZIP files
- -O /media/sf_exchange/sentineldata: folder where downloaded Sentinel data will be stored
- -R /media/sf_exchange/sentinel_data/failed.txt: file listing products which failed to download
- -F ‘query’: this is the full text query you can copy/paste from the ESA SciHUB website
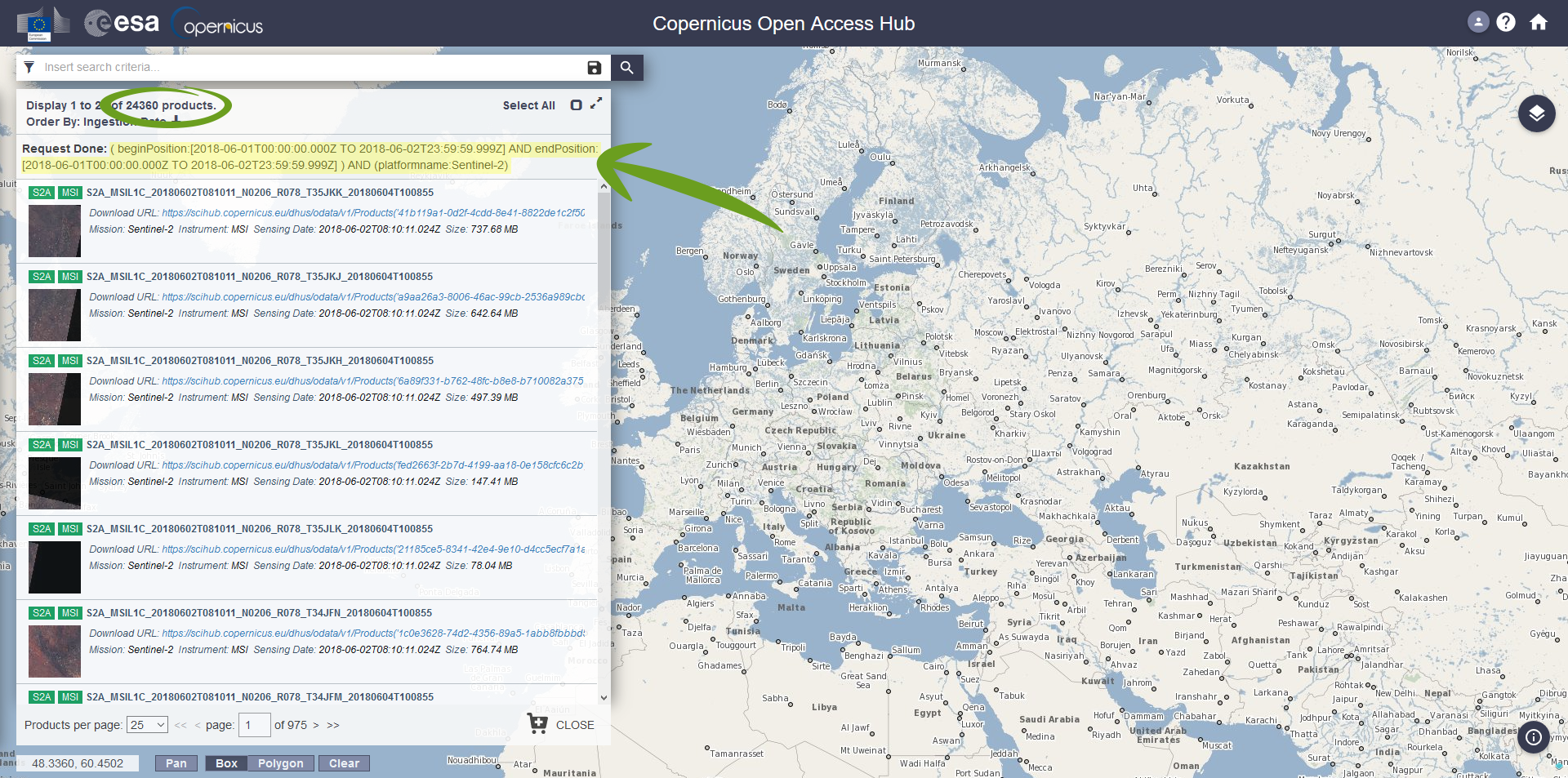
If you use our VM, you just need to replace your username -u user123 and password -p password123 with your own login information. Actually, you only need to integrate your search query in -F ‘query’ as a last step. Thus, visit the ESA Sci-HUB portal and set your data query to your liking as shown in Perform a Search section. In the upper part of the result window, the query can be obtained in text form. This is an example of a query giving all Sentinel 2 products sensed between the first and second of June 2018 globally:
You will want to mark and copy the whole query expression and paste it between the simple quotes of the -F ‘query’ option, like so:
-F ' ( beginPosition:[2018-06-01T00:00:00.000Z TO 2018-06-02T23:59:59.999Z] AND endPosition:[2018-06-01T00:00:00.000Z TO 2018-06-02T23:59:59.999Z] ) AND (platformname:Sentinel-2) '
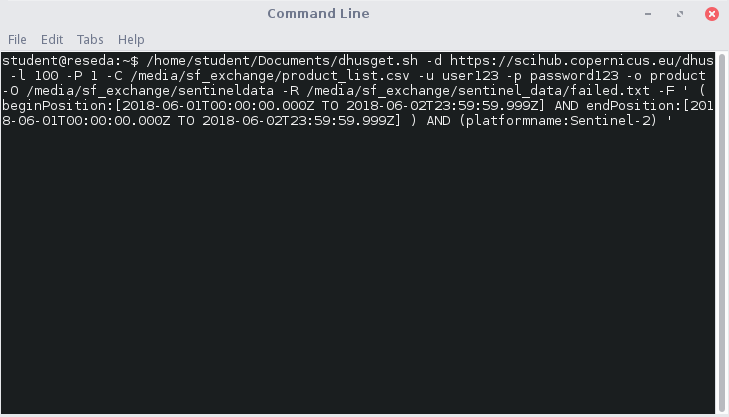
Open a terminal (taskbar in our VM) and type in the complete query, e.g.:
By pressing enter it will start downloading up to 100 scenes at once. Previously downloaded scenes will not be downloaded again, and if the internet connection is lost, the download will continue at that point when restarting the script. When a scene is fully loaded, the MD5 integrity will be checked, telling you whether the file is complete or not. If not, the scene is written to a text file to notify you -R /media/sf_exchange/sentinel_data/failed.txt.
By doing a search query on the ESA SciHUB Website, you can also see the resulting products available (which is the insane amount of 24,360 in our example – you probably do not need that many). However, for some reason ESA capped the maximum amount of scenes you can query/download at once at 100. Because of that, you will only download the search results 1-100. In order to download search results 101-200 you have to change the result page with the option -P 2, for the search results 201-300 set this option to -P 3, and so on. If required, this could also be automated, either by a shell script or in R.